This set of Molecular Biology Questions and Answers for Aptitude test focuses on “Overview of Protein Synthesis – 2”.
1. Eukaryotic ribosome is recruited to the mRNA by _____________
a) Randomly
b) Shine – Dalgarno sequence
c) 5’ capping
d) 3’ tailing
View Answer
Explanation: Prokaryotic ribosome is recruited to the mRNA by the Shine – Dalgarno sequence. In the case of eukaryotes ribosome is recruited to the mRNA by the 5’ cap.
2. After bounding to the mRNA eukaryotic ribosome scans the entire mRNA until it encounters a start codon.
a) True
b) False
View Answer
Explanation: The eukaryotic ribosome is recruited to the mRNA by the 5’ cap. The guanine residue of the 5’ cap is connected to the 5’ end of the mRNA through three phosphate groups that recognizes and recruits the ribosome. Once bound to the mRNA, the ribosome moves in a 5’→3’ direction until it encounters a 5’…….AUG…….3’ start codon by a process called scanning.
3. The end of all tRNAs is ___________
a) 5’ ACC 3’
b) 5’ CCA 3’
c) 3’ CAC 5’
d) 3’ GAG 5’
View Answer
Explanation: All tRNAs end at the 3’ terminus with the sequence 5’ CCA 3’. This is the site that is attached to the cognate amino acid by the enzyme aminoacyl tRNA synthetase.
4. How many unusual bases are observed in a tRNA molecule?
a) 2
b) 3
c) 4
d) 5
View Answer
Explanation: Five unusual bases, produced by enzymatic modification of the usual bases, may be observed in a tRNA molecule. These are pseudouridine, dihyrdouridine, hypoxanthine, thymine and methylguanine. Of these five, unusual bases pseudouridine and dihyrdouridine are most commonly found in the tRNA molecules.
5. In pseudouridine the attachment of Uracil base to ribose through ___________ of uracil.
a) N at position 3
b) C at position 6
c) C at position 5
d) N at position 1
View Answer
Explanation: Pseudouridine is produced by the enzymatic modification of the regular base uridine through isomerization. In the modification process of uridine to pseudouridine the attachment of base to the ribose sugar is switched from nitrogen at ring position 1 to the carbon at ring position 5.
6. Dihydrouridine is the result of oxidation of the uracil base.
a) True
b) False
View Answer
Explanation: Dihydrouridine is a derivative of uridine formed by enzymatic reduction. The process of reduction forms the double bond between the carbons at positions 5 and 6.
7. How many loops are seen in the tRNA?
a) 1
b) 2
c) 3
d) 4
View Answer
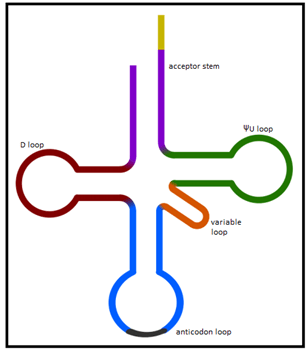
Explanation: Four different loops are observable in the tRNA 3˚ motif. They are the ΨU loop, D loop, variable loop and the anticodon loop. The ΨU loop is named because of the presence of the pseudouridine base. The D loop contains the dihyrdouridine base. The anticodon loop contains the anticodon and the variable loop is named so because its length is variable.
8. How many double helix regions are observed in a tRNA molecule?
a) 4
b) 3
c) 2
d) 1
View Answer
Explanation: The tRNA molecules consist of 4 double helix regions. They are the acceptor stem and the stem of the three loops, the ΨU loop, D loop and the anticodon loop.
9. The variable loop is found between the ___________
a) Acceptor stem and ΨU loop
b) Acceptor stem and D loop
c) Anticodon loop and ΨU loop
d) Anticodon loop and D loop
View Answer
Explanation: The variable loop sits between the anticodon loop and ΨU loop. Also as the name implies the size of the variable loop varies from 3 to 21 bases.
10. The 3 dimension structure of an adaptor molecule of translation is revealed to be ___________
a) Y shaped
b) L shaped
c) K shaped
d) X shaped
View Answer
Explanation: The adaptor molecule of translation is the tRNA molecule which acts as the bridge between the mRNA code and the respective amino acid. X-ray crystallography reveals an L-shaped tertiary structure in which the terminus of the acceptor stem is at one end of the molecule and the anticodon loop is about 70 Å away at the other end.
11. Which of the following does not take part in stabilizing the cloverleaf model of the tRNA?
a) Base stacking
b) Base and sugar-phosphate backbone interaction
c) Ionic bond
d) Hydrogen bond
View Answer
Explanation: Three types of interactions stabilize the L-shaped structure of the tRNA. The first is hydrogen bonds between the bases to form the helical parts of the tertiary structure of the tRNA molecule. Second the interaction between the bases and the respective sugar phosphate backbone. Finally, the additional stabilization is provided by the base stacking between the two extended regions of base pairing.
Sanfoundry Global Education & Learning Series – Molecular Biology.
To practice all areas of Molecular Biology, here is complete set of 1000+ Multiple Choice Questions and Answers.
If you find a mistake in question / option / answer, kindly take a screenshot and email to [email protected]
- Check Biotechnology Books
- Check Molecular Biology Books
- Apply for Biotechnology Internship
- Practice Biotechnology MCQs
